Mass finder chemistry6/2/2023 
The standard atomic weights of carbon, hydrogen, and oxygen are approximately 12.011 , 1.008 , and 15.999 g/mol respectively. Let us see approximately how many grams are in 2.3 moles of table sugar with a chemical formula of C 12H 22O 11. A given polymer sample is said to be made up of a mixture of macromolecules with a certain molar mass distribution. 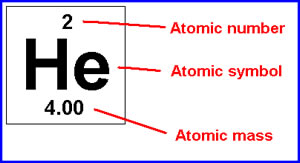
In linear polymers not every polymer chain consists of the same amount of repeating units. In physics, molar mass is usually defined in kilograms per mole (kg/mol) because the base SI unit of mass is the kilogram. In chemistry, the unit of molar mass is g/mol due to chemical utility. Molar mass is different from Molecular mass which is the mass of one molecule. Thus the molar mass is the mass of 6.023x10 23 particles of the substance. A mole of a substance is defined to be approximately 6.023x10 23 ( see Avogadro's number) of particles of the substance. 
The molar mass of a chemical substance may be computed from the standard atomic weights listed for the elements on a standard periodic table. For example, if an equal number of molecules of two substances are needed for a reaction but the molar mass of one substance is twice that of the other, twice as many grams will be needed of that substance to give the same number of molecules. This allows for using the appropriate number of molecules of a substance regardless of the mass. Its primary purpose is as a conversion factor between the number of grams of a pure substance, which can be measured directly, and the number of moles of that substance, which has greater chemical significance. It is commonly used in stoichiometric calculations of bulk substances in chemistry. Version 2.0 includes the ability to obtain the expanded uncertainty (k=2) of the calculated result based on Monte Carlo simulation (100,000 trials) along with more modification options.Molar mass is the mass of one mole of a chemical element or chemical compound. A useful feature is the ability to add a desired mass to a specific residue enabling the identification of fragment peak profiles which match experimental data. The Residue Codes menu option lists the valid single-letter codes for the residues including some modified residues. The total mass of the sub-unit will be displayed including selected additions such as pyroglutamation, glycosylation and disulfide bonds. The sequence mass is the sum of the product of the accumulated totals (plus water) by the elemental masses obtained from the following link:Ĭalculations may be performed on multi-subunit proteins by separating sequences with an ampersand (&). 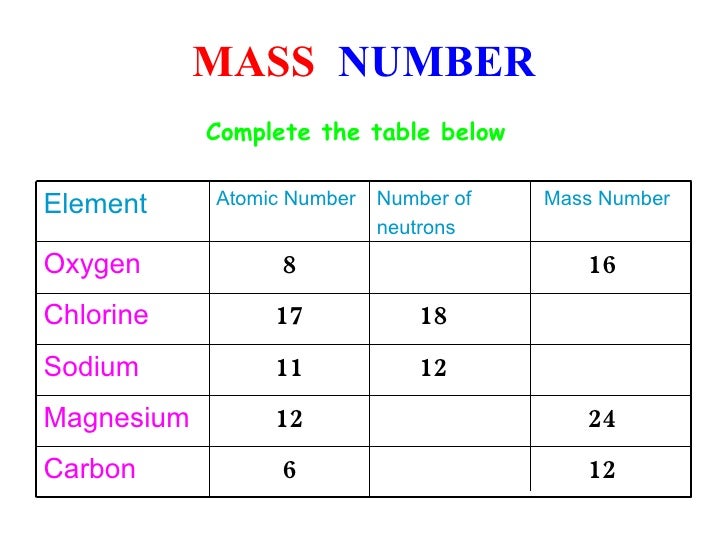
Sequence Masses are calculated based on the accumulated occurrences of all the elements within each residue contained in the input sequence according to the residue formula displayed in the Residue Formula menu option. z-ETD is the fragmentation associated with ETD or ECD as shown in Syka et al. 59(21) 2621-2625 (1987)) which is relevant to higher-energy fragmentation associated with CID. z-CID is the structure from Johnson et al. Two forms of the z product ions are displayed. The option exists to select monoisotopic results which will also generate y, b, c, and z product ions. The NIST Mass and Fragment Calculator is a program written in Visual Basic which calculates the mass of an input peptide or protein sequence along with m/z ions corresponding to 1+, 2+, and 3+ charge states when selecting average mass results.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
